-Search query
-Search result
Showing 1 - 50 of 143 items for (author: lee & dj)

EMDB-43658: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43659: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43660: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)
EMDB-40242: 
BG505 MD39 SOSIP in complex with Rh.NJ85 wk12 gp120GH, N611/FP, and base epitope polyclonal antibodies
Method: single particle / : Lee WH, Ozorowski G, Torres JL, Ward AB

EMDB-40243: 
BG505 MD39 SOSIP in complex with Rh.NJ86 wk12 V1V3, C3V5, N611/FP, and base epitope polyclonal antibodies
Method: single particle / : Lee WH, Ozorowski G, Torres JL, Ward AB

EMDB-40244: 
BG505 MD39 SOSIP in complex with Rh.NJ76 wk12 C3V5, N611/FP, and base epitope polyclonal antibodies
Method: single particle / : Lee WH, Ozorowski G, Torres JL, Ward AB
EMDB-40252: 
BG505 MD39 SOSIP in complex with Rh.NK04 wk12 gp120 glycan hole and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40254: 
BG505 MD39 SOSIP in complex with Rh.NJ75 wk12 gp120 glycan hole and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40255: 
BG505 MD39 SOSIP in complex with Rh.NJ87 wk12 C3V5 and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40256: 
BG505 MD39 SOSIP in complex with Rh.NJ84 wk12 V1V3, gp120 glycan hole and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40257: 
BG505 MD39 SOSIP in complex with Rh.NJ77 wk12 V1V3, C3V5, N611/FP and base epitope polyclonal
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-28910: 
Glycan-Base ConC Env Trimer
Method: single particle / : Olia AS, Kwong PD

EMDB-29725: 
Vaccine-elicited human antibody 2C06 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Morano NC, Shapiro L, Kwong PD

EMDB-29731: 
Vaccine-elicited human antibody 2C09 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD

EMDB-27876: 
E. coli 50S ribosome bound to tiamulin and VS1
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27877: 
E. coli 50S ribosome bound to tiamulin and azithromycin
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27878: 
E. coli 50S ribosome bound to compound streptogramin A analog 3336
Method: single particle / : Pellegrino J, Lee DJ, Seiple IB, Fraser JS

EMDB-27867: 
E. coli 50S ribosome bound to D-linker solithromycin conjugate
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27868: 
E. coli 50S ribosome bound to L-linker solithromycin conjugate
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27869: 
E. coli 50S ribosome bound to solithromycin and VM1
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27852: 
E. coli 50S ribosome bound to compound streptogramin A analog 3142
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27854: 
E. coli 50S ribosome bound to compound streptogramin analogs SA1 and SB1
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27855: 
E. coli 50S ribosome bound to compound streptogramin analog SAB001
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27857: 
E. coli 50S ribosome bound to compound SAB002
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27858: 
E. coli 50S ribosome bound to compound streptogramin A analog 3146
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27879: 
E. coli 50S ribosome bound to antibiotic analog SLC09
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27880: 
E. coli 50S ribosome bound to antibiotic analog SLC17
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27881: 
E. coli 50S ribosome bound to antibiotic analog SLC21
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27882: 
E. coli 50S ribosome bound to antibiotic analog SLC26
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27883: 
E. coli 50S ribosome bound to antibiotic analog SLC30
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27884: 
E. coli 50S ribosome bound to antibiotic analog SLC31
Method: single particle / : Pellegrino J, Lee DJ, Fraser JS, Seiple IB

EMDB-27943: 
9H2 Fab-poliovirus 1 complex
Method: single particle / : Charnesky AJ
EMDB-27947: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

EMDB-27948: 
9H2 Fab-poliovirus 2 complex
Method: single particle / : Charnesky AJ
EMDB-27949: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ
EMDB-27950: 
9H2 Fab-Sabin poliovirus 2 complex
Method: single particle / : Charnesky AJ
EMDB-27951: 
9H2 Fab-Sabin poliovirus 1 complex
Method: single particle / : Charnesky AJ

EMDB-16227: 
Hexameric human IgG3 Fc complex
Method: subtomogram averaging / : Abendstein L, Sharp TH

EMDB-16241: 
Subtomogram average of a IgG3-C1-C4b complex on a lipid bilayer
Method: subtomogram averaging / : Abendstein L, Sharp TH

EMDB-16250: 
Subtomogram average of a complex of IgG3-C1-C4b after focussed refinement around the C4b region
Method: subtomogram averaging / : Abendstein L, Sharp TH
EMDB-16251: 
Subtomogram average of a complex of IgG3-C1-C4b after focussed refinement around the C1 region
Method: subtomogram averaging / : Abendstein L, Sharp TH

EMDB-14866: 
VelcroVax tandem HBcAg with SUMO-Affimer inserted at MIR (T=4 VLP)
Method: single particle / : Kingston NJ, Grehan K, Snowden JS, Alzahrani J, Ranson NA, Rowlands DJ, Stonehouse NJ

EMDB-14868: 
VelcroVax tandem HBcAg with SUMO-Affimer inserted at MIR (T=3* VLP)
Method: single particle / : Kingston NJ, Grehan K, Snowden JS, Alzahrani J, Ranson NA, Rowlands DJ, Stonehouse NJ
EMDB-27898: 
Cryo-EM structure of human glycerol-3-phosphate acyltransferase 1 (GPAT1) in complex with 2-oxohexadecyl-CoA
Method: single particle / : Johnson ZL, Wasilko DJ, Ammirati M, Chang JS, Han S, Wu H
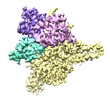
EMDB-27899: 
Cryo-EM structure of human glycerol-3-phosphate acyltransferase 1 (GPAT1) in complex with CoA and palmitoyl-LPA
Method: single particle / : Wasilko DJ, Johnson ZL, Ammirati M, Chang JS, Han S, Wu H

EMDB-15725: 
Poliovirus type 3 (strain Saukett) stabilised virus-like particle (PV3 SC8) in complex with GSH and GPP3
Method: single particle / : Bahar MW, Fry EE, Stuart DI
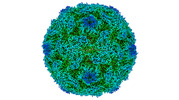
EMDB-15726: 
Poliovirus type 3 (strain Saukett) stabilised virus-like particle (PV3 SC8) in complex with GSH and Pleconaril
Method: single particle / : Bahar MW, Fry EE, Stuart DI
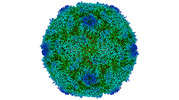
EMDB-15727: 
Poliovirus type 2 (strain MEF-1) virus-like particle in complex with capsid binder compound 17
Method: single particle / : Bahar MW, Fry EE, Stuart DI

EMDB-15543: 
Poliovirus type 3 (strain Saukett) stabilised virus-like particle (PV3 SC8).
Method: single particle / : Bahar MW, Fry EE, Stuart DI

EMDB-27610: 
BG505 MD39 SOSIP in complex with Rh.NJ82 wk13 N611 and base epitope pAbs
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
